 Yourself with Aligner's most important features. You to try the out all functions in CodonCode Aligner.Īfter downloading, we suggest that you watch the introduction First-time users get an automatic 30-day trial that allows 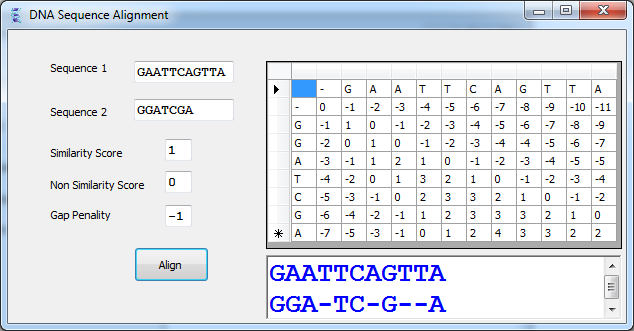
The demo version is a fully functional trace Assemble bacterial genomes and small NGS sequencing projectsĬodonCode Aligner lets you take full advantage of modern algorithms for base calling and sequence assembly, withoutįorcing you to learn complicated command line options or Perl programming.ĭemo version of CodonCode Aligner.Define regions of interest ("features"), for.Pland and visualize restriction cloning steps.Analyze methylated sequences & view raw trace data.Find enzymes for RFLP analysis & create virtual gels.See difference tables and protein translations.Run Phred and Phrap directly from Aligner projects.Generate phylogenetic trees and restriction maps.Quickly navigate between user-defined features.Analyze heterozygous insertions and deletions.Assemble or align in groups, based on sample names.Align contigs to each other while keeping links to the traces.Import sequences from ABI, SCF, and FASTQ files.GenBankĪssumes that the submitter has received any necessary informed consentĪuthorizations required prior to submitting sequences. If you are submitting human sequences to GenBank, do not include anyĭata that could reveal the personal identity of the source. As soon as it is available, please send the full publication data-all authors, title, journal, volume, pages and date-to the following address: Privacy In order to prevent the delay in the appearance of published sequence data, we urge authors to inform us of the appearance of the published data. However, if the accession number or sequence data appears in print or online prior to the specified date, your sequence will be released. This is why we compiled on a list of software and tools which supports you processing and interpreting your experimental data, be that next-generation sequencing, microarray or mass spectrometry. GenBank will, upon request, withhold release of new submissions for a specified period of time. Some authors are concerned that the appearance of their data in GenBank prior to publication will compromise their work. More details about this process can be found on the NLM GenBank and SRA Data Processing. On rare occasions, data may be removed from public view. Following submission, data are subject to automated and manual processing to ensure data integrity and quality and are subsequently made available to the public. The most important source of new data for GenBank is direct submissions from a variety of individuals, including researchers, using one of our submission tools. Permission concerning the use, copying, or distribution of the NCBI is not in a position to assess the validity of suchĬlaims, and therefore cannot provide comment or unrestricted Property rights in all or a portion of the data they have Some submitters may claim patent, copyright, or other intellectual Restrictions on the use or distribution of the GenBank data. Within the scientific community to the most up-to-date andĬomprehensive DNA sequence information. The GenBank database is designed to provide and encourage access The ASN.1 and flatfile formats are available at NCBI's anonymous FTP.Search, link, and download sequences programatically using NCBI e-utilities. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
